Research
Current research in the Videvall Lab — Uppsala University
Diet-microbiome associations in the Scandinavian wolverine (led by Charlotte)
Impacts of feralization on the horse microbiome (led by Madeleine)
Effects of captivity on the European bison gut microbiome (led by Nils)
Microbiome, diet, and metabolism in hybrid flycatchers (led by Ester and Qianrui)
Flycatcher cross-fostering experiment (led by Ester)
Sand lizard cloacal microbiomes (led by Charlotte)
Role of temperature and diet on toad tadpole gut microbiomes (led by Gabriel)
Genome-microbiome interactions in Hawaiian feral chickens (led by Matias)
Temporal hologenomics of historical gut microbiomes pilot project (led by Elin)
Microbiota and AMR transmission in chironomids (postdoc position open, Lab assistant: Ramin)
Early-life gut microbiomes associated with growth and health in reindeer calves (upcoming project)
Feral horses in northern Spain, photo by Elin Videvall
Pied flycatcher, photo by Hans Ollonen
Chironomid larva, photo by Frank Fox
European bison, photo by Charles J. Sharp
Wolverine, photo by Susanne Nilsson
Reindeer, photo by Ingebjørg Nymo
Sand lizard, photo by George Chernilevsky
Feral chicken, photo by Richard N Horne
Current research collaborators
Dr. Mette Lillie, Uppsala University
Prof. Anna Qvarnström, Uppsala University
Dr. Carl-Gustaf Thulin, SLU, Uppsala
Dr. Germán Orizaola, University of Oviedo
Prof. Dominic Wright, Linköping University
Dr. Bruno Carreira, University of Lisbon
Dr. Jamie Muriel, University of Córdoba
Prof. Frank Johansson, Uppsala University
Dr. Richard Svanbäck, Uppsala University
Dr. David Canal, National Museum of Natural Sciences, Madrid
Dr. Robert Ekblom, Swedish Environmental Protection Agency
Dr. Göran Spong, SLU, Umeå
Dr. Josué Martínez-de la Puente, Doñana Biological Station
Prof. Anna Skarin, SLU, Uppsala
Dr. Peter Halvarsson, SLU, Uppsala
Elin’s postdoctoral research — Brown University
Giraffe cross-species diet-microbiome associations (Videvall et al. 2025 Glob. Ecol. Conserv.)
Diet and microbiota changes in giraffes in response to human barriers
Diet-microbiome associations in Ugandan giraffes
PacBio shotgun metagenomics of giraffe microbiomes
Phylosymbiosis of mammalian microbiomes
Elin’s postdoctoral research — Smithsonian
Avian malaria genomics - current and historical spread of Plasmodium relictum throughout the Hawaiian islands (delayed due to Covid-19)
Genomics of Hawaiʻi ‘amakihi through time (historical DNA from museum samples) (delayed due to Covid-19)
Microbiomes of Hawaiʻi ‘amakihi with malaria infection (advisory role - Navine et al. 2022 Mol Ecol)
Microbiome phylosymbiosis and diet of Hawaiian honeycreepers (advisory role - Costantini et al. Ecol Ecol)
Transcriptomics of the Hawaiian strain of Plasmodium relictum (Videvall et al. 2021 Ecol & Evol)
Transcriptomics of Hawaiʻi ‘amakihi during malaria infection (collaboration - Paxton et al. 2023 J Hered)
Transcriptomics of Culex mosquitoes to Plasmodium infection (collaboration - Ferreira et al. 2022 Malaria J).
Istmobiome - Host-microbe associations using natural experiments in marine environments (Leray et al. 2021 PLOS Biol)
Elin’s PhD research — Lund University
Evolutionary genomics of host-microbe interactions
Elin’s PhD thesis included ten papers in two different projects:
1. Genomic interactions between malaria parasites and avian hosts
2. The gut microbiome's influence on avian fitness and survival
Genomic interactions between malaria parasites and avian hosts
We used dual RNA-sequencing of birds infected with malaria parasites. Because we sequence the blood, this allowed us to analyse not only the transcriptome of the host, but also the transcriptome of the parasite simultaneously. Resulting in an unprecedented opportunity to evaluate real-time interactions between hosts and parasites.
Publications in this project:
Videvall et al. (2015) The Avian Transcriptome Response to Malaria Infection. Molecular Biology and Evolution.
Bensch et al. (2016) The genome of Haemoproteus tartakovskyi and its relationship to human malaria parasites. Genome Biology and Evolution.
Videvall et al. (2017) The transcriptome of the avian malaria parasite Plasmodium ashfordi displays host-specific gene expression. Molecular Ecology.
Hellgren et al. (2017) De novo synthesis of thiamine (vitamin B1) is the ancestral state in Plasmodium parasites - evidence from avian haemosporidians. Parasitology.
Videvall. (2018) Plasmodium parasites of birds have the most AT-rich genes of eukaryotes. Microbial Genomics.
Videvall et al. (2020) Host transcriptional responses to high- and low-virulent avian malaria parasites. The American Naturalist.
The gut microbiome's influence on avian fitness and survival
Ostrich chicks can experience dramatic mortality rates due to disease. The development of a healthy gut microbiome seems to be a crucial step in the first three months of an ostrich.
We used 16S rRNA Illumina sequencing to characterize the gut microbiome over time and associate its various components with growth and survival.
Publications in this project:
Videvall et al. (2017) Direct PCR offers a fast and reliable alternative to conventional DNA isolation methods for gut microbiomes. mSystems.
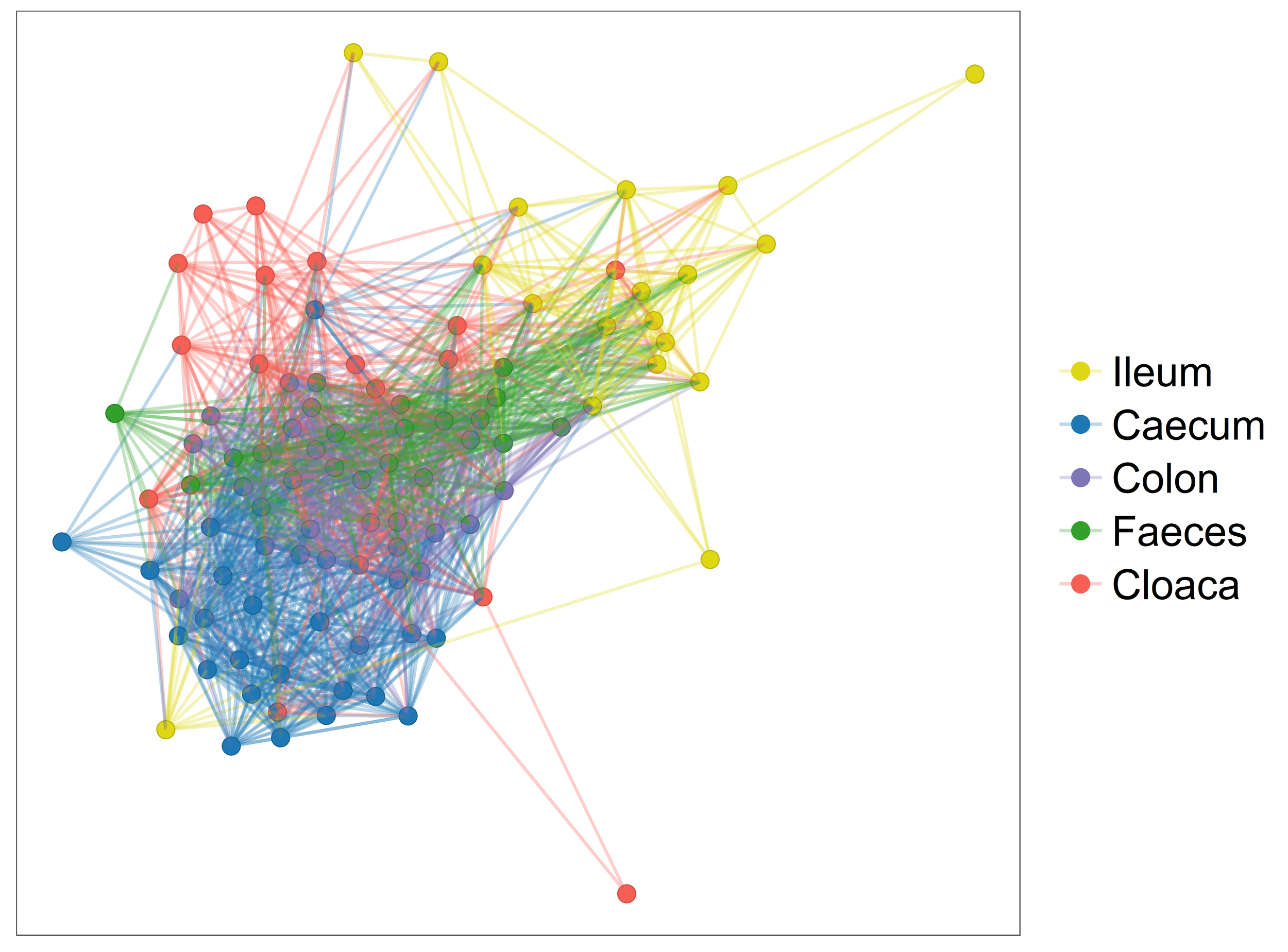
Videvall et al. (2017) Measuring the gut microbiome in birds: comparison of faecal and cloacal sampling. Molecular Ecology Resources.
Videvall et al. (2019) The colonization of gut microbiota during development and its association with growth in ostriches. Molecular Ecology.
Videvall et al. (2020) Early-life gut dysbiosis linked to mass mortality in ostriches. Microbiome.
Videvall et al. (2023) Coprophagy rapidly matures juvenile gut microbiota in a precocial bird. Evolution Letters.
Previous research & collaborations
Chernobyl soil microbiomes (Videvall et al. 2022)
Preen gland microbiome (Videvall et al. 2021)
Sex-specific gene expression in birds using RNA-sequencing (Sigeman et al. 2018)
Transcriptome differences between urban and rural birds - (Watson et al. 2017)
Transcriptomics of outcrossing Arabidopsis lyrata - (Videvall et al. 2016)
Transcriptomics of the hessian fly - (Andersson et al. 2014)
Molecular identification of blood meals in biting midges - (Videvall et al. 2013)
Monitoring butterflies using grid-based sampling - (Videvall et al. 2016)